RACIAL
INFLUENCE ON THE PREVALENCE OF PROSTATE CARCINOMA IN BRAZILIAN VOLUNTEERS
(
Download pdf )
EDSON L. PASCHOALIN, ANTONIO C.P. MARTINS, MÔNICA PASTORELLO, KIYOKO A. SÂNDIS, LEA M.Z. MACIEL, WILSON A. SILVA JR., MARCOS A. ZAGO, JOSÉ BESSA JR.
Ribeirão Preto General Hospital, School of Medicine, USP, São Paulo, Brazil
ABSTRACT
Purpose:
To investigate the prevalence of prostate carcinoma in a sample of volunteers
known to have a large proportion of Bantu African ancestors, and the performance
of total PSA (tPSA), PSA density (PSAD) and free-to-total PSA ratio (f/tPSA)
on the diagnosis.
Materials and Methods: A total of 473 volunteers
(range: 40 - 79 years) were screened for prostate carcinoma. Those with
tPSA >2 ng/ml and/or abnormal digital rectal examination were submitted
to a transrectal ultrasound-directed biopsy (10 cores). The volunteers
were classified as White, Mulatto or Black according to physical characteristics
and to ancestors race reference. The following variable number of tandem
repeats (VNTR) were analyzed in the blood of 120 volunteers without cancer
and in 27 patients with prostate cancer: D4S43, PAH, F13A1, APOB and vW-1.
Results: The biopsies performed in 121 volunteers
revealed cancer in 27 (5.7% of 473). The proportions of cancer in White,
Mulatto and Black were respectively: 0.6% (1/148), 6.7% (6/90) and 8.5%
(20/235) (p = 0.006). The VNTRs analysis revealed heterogeneity in White,
Mulatto and Black anthropologic phenotypes with the following admixture
of Caucasian, African and Amerindian gene lineages: 67.5 ± 8%,
20.8 ± 8%, 11.7 ± 7%; 54.8 ± 9%, 36.3 ± 5%,
8.9 ± 7%; and, 45.3 ± 3%, 45.9 ± 4%, 8.8 ±
7%. Such a mixture was 50.5 ± 9%, 49 ± 8% and 0.5 ±
4% in volunteers bearing cancer, and 59.1 ± 7%, 31.7 ± 8%
and 9.2 ± 5% in those without cancer. The sensitivity and specificity
of tPSA at cut-off levels of 2, 2.5 and 4 ng/ml for volunteers with tPSA
£ 10 ng/ml were respectively: 100% and 6,6%, 100% and 36,6%, 69,2%
and 62,2%. PSAD at a cut-off level of 0.08 or 0.10, and f/tPSA at a cut-off
level of 20% were able to increase significantly tPSA specificity without
loss on sensitivity.
Conclusions: The tumor prevalence was higher
in Non-White than in White phenotype. The association of tPSA at a cut-off
level of 2.5 ng/ml with a PSAD of 0.08 or a f/tPSA of 20% for biopsy indication
deserves further investigations as an alternative to tPSA cut-off level
of 4 ng/ml.
Key
words: prostatic neoplasms; prevalence; race; prostate-specific
antigen
Int Braz J Urol. 2003; 29: 300-5
INTRODUCTION
The
prevalence of prostate adenocarcinoma is 50% higher in North-American
Afro-American than in Caucasians, and it can be 3 or 4 times higher when
compared to Chinese and Japanese (1). At the moment of diagnosis in Afro-Americans,
the stage is more advanced, and for this reason the earlier beginning
of screening has been advocated in Afro-American men (2).
The definition for the normality range of
the prostate specific antigen (PSA), published in 1986 by Myrtle et al.
(3), is largely accepted. This group from Hybritech, using Tanden-R assay,
established the interval between 0 and 4 ng/ml as the normality range.
One among 3 men with a level superior to that will show carcinoma in prostate
biopsy (4). However, there are reports proposing changes in this range,
which could vary according to race and age (1).
Gann et al. (5) showed that men with total
PSA (tPSA) between 2 and 4 ng/ml are 12-fold more likely to develop prostate
cancer compared to others with tPSA below 1 ng/ml, when followed during
10 years. There are studies showing that in the tPSA range between 2.5
and 4 ng/ml the prevalence of cancer reaches 24%, and that is why many
centers started to recommend this cut-off level for indication of biopsy
(6).
In Brazil, studies on screening for prostate
adenocarcinoma are scarce. The existing data present similarity in sensitivity
and specificity of tPSA in a cut-off of 4 ng/ml, for Southeastern population,
as observed in North American population, but there are no reports of
investigation using lower cut-off levels. On the other hand, 3 studies
with Southeastern population involving 5,313 volunteers were not able
to convincingly demonstrate a higher prevalence of prostate cancer in
Afro-Brazilians than in Caucasians (7-9). It must be stressed that about
70% of Brazilian black population originate from Angola, Congo and Mozambique
where the Bantu haplotype is predominant (10). The study of hemoglobin
b genes confirms that in Brazil 73% of the haplotypes are Bantu type,
with Senegal haplotype being practically inexistent (11). These data reveal
that Afro-Brazilian are genetically distinct from North American black
population, where the haplotype Benin predominates (59%) with equivalent
frequencies of Bantu and Senegal haplotypes (12). Additionally, studies
based in polymorphism of nuclear and mitochondrial DNA stress the role
of the intense process of miscegenation undergone by Afro-Brazilian population
(urban or isolated – remainders of the quilombos) (13). The African
component in black populations can range from 46 to 67% in “rural”
areas, but these data do not have universal validity for urban populations
(14). Obviously, the ethnic admixture is not restricted to Afro-Brazilian
descendants, but it is extended to Caucasians. Such data warrant the performance
of screenings in our environment aimed to racial prevalence of prostate
tumor and a better characterization of PSA.
MATERIALS AND METHODS
The
target population for screening consisted of 473 volunteers with ages
ranging from 40 to 79 years from the city of Ipirá, Bahia. All
of them underwent a digital rectal examination, blood collection for dosage
of tPSA and free PSA (DPC – Immulite test)Ò and DNA extraction
for genetic race analysis. Volunteers with tPSA equal or superior to 2
ng/ml or digital rectal examination suspected of prostate cancer underwent
transrectal ultrasound-directed prostate biopsy (at which time the prostate
volume was measured) collecting 10 fragments from the peripheral zone
(15). These fragments were conserved in 10% formalin solution until their
processing for histological examination following hematoxylin-eosin staining.
Volunteers were classified in White, Mulatto
or Black according to anthropological criteria that considered not only
the skin color aspect, but the ancestors’ racial reference up to
3rd degree (great-grandparents).
Racial genetic diversity was studied through
variable number of tandem repeats (VNTR) in the DNA of 40 volunteers randomly
chosen from each of the 3 racial groups defined by the anthropological
criteria, as well as all bearers of neoplasia. DNA extraction was performed
by a standardized technique, and its amplification (PCR) was made with
35 cycles in “Perkin-Elmer-CetusÒ” thermocycler employing
the following starters: APO B, vW-I Factor, D4S43, PAH and 13A1 Factor
(13). The different alleles were recognized through electrophoretic migration
in 4% polycrilamide gel after marking PCR products with flurochrome. Differences
in molecular weight (or number of bases) were determined by comparison
with migration of the “Gene Scan 2500 ROXÒ Kit” standard
using the equipment’s “ABIPrism 377Ò” software.
The organization of the database with frequency of alleles for comparison
between groups was done by the GDA software (16). That computerized VNTR
database was employed as a reference, with data on descendants with European
origin (Portuguese, Spanish, Italian and German – urban region of
Ribeirão Preto), Africans from Congo and Cameroons (origin: Lubumbashi
and Yaoundé) and Amerindian from isolated tribes of Pará
(Arara, Wayana-Apalai, Wayampi, Yanoman and Kayapo) (13). The estimate
of ethnic admixture was calculated by the ADMIX 3 software (17).
The variables related to cancer prevalence,
PSA test and other characteristics of the sample were performed by the
Graph PrismÒ software version 3.0. Classificatory attributes were
analyzed by Fisher’s exact test or c2. Continuous variables of normal
distribution were compared by unpaired bi-caudal “t” test
or variance analysis, and those which did not pass the normality test
were assessed by non parametric test. The significance level was fixed
in 5%.
RESULTS
Of
the 473 volunteers, 148 (31.3%) were White, 90 (19%) Mulatto and 235 (49.7%)
Black according to the anthropological criteria. Respective mean age in
these groups was: 56.8 ± 9.5 years, 54.7 ± 10.7 years and
57.9 ± 9.5 years. Simultaneous comparison between the groups by
variance analysis showed p = 0.03. Tukey test showed similarity between
White and Black, as well as between White and Mulatto groups, but not
between Mulatto and Black groups. The comparison of mean age of Whites
and non-Whites by the t-test showed no difference (p = 0.8).
Biopsies were indicated and performed in
121 volunteers. The prevalence of prostate adenocarcinoma for a tPSA cut-off
level of 2 or 2.5 ng/ml was 5.7%, or 27/473 men, with ages ranging from
40 to 79 years, and 7.9% or 27/341 volunteers with ages between 50 and
79 years. If a tPSA cut-off level of 4 ng/ml was considered the prevalence
would be 23/473 (4.8%) between 40 and 79 years, and 23/341 (6.7%) from
50 to 79 years.
Tumor occurred in 1 White, 6 Mulattos and
20 Blacks. Simultaneous comparison of cancer prevalence in the 3 groups,
according to the anthropological criteria, showed a high significance
(p = 0.005). The comparison between groups showed the following results:
White versus Black: p = 0.008, White versus Mulatto: p = 0.01, White versus
non-White: p = 0.009 and Mulatto versus Black: p = 0.65.
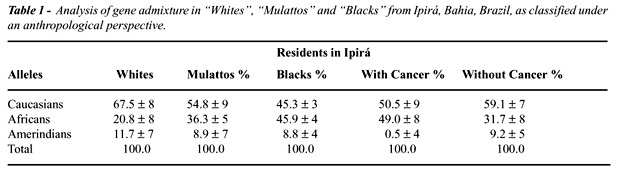
The mean values of admixture rate of African,
Caucasian and Amerindian genes, in the samples classified by anthropological
criteria as White, Mulatto or Black are exposed in Table-1. The gene admixture
in volunteers with or without cancer regardless of anthropological phenotype
is presented in this Table as well.
Tumor ratios in tPSA ranges from 2.1 to
4 ng/ml, from 2.5 to 4 ng/ml, from 4 to 10 ng/ml and >10 ng/ml were
respectively: 6/62 (9.6%), 6/43 (13.9%), 7/35 (20%) and 14/30 (77.7%).
The performance of the tPSA test in 3 cut-off levels in volunteers with
tPSA £ 10 ng/ml is exposed in Table-2.
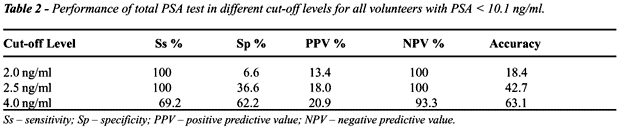
Table-3 shows the performance of digital
rectal examination as well as a simulation of influence, on the performance
of tPSA, of free/total PSA ratio and of PSA density in cases they were
used. For simulation, the selection of cut-off levels of free/total PSA
ratio and PSA density was made in such a way to maintain the sensitivity
of total PSA according to the cut-off level.

DISCUSSION
The
prevalence of prostate cancer in volunteers from Bahia community (Northeastern)
under study seems equivalent to those found in the State of São
Paulo since those ranged from 1.3% to 3.2% (7-9,18). It must be stressed
that in Northeastern volunteers the prevalence would be 4.8% if a tPSA
cut-off level 4 ng/ml was employed, and that the performance of biopsies
with collection of 10 peripheral fragments can lead to a 35% addition
in cancer diagnosis compared to the sextant biopsy (15). In developed
countries, the prevalence of prostate cancer in screenings has ranged
from 1% to 6% in the age range from 50 to 75 years (1,4,6), which appears
to indicate a similarity with our population.
Tumor prevalence in Blacks and Mulattos,
alone or jointly, was statistically superior to that found in Whites from
Ipirá. However the prevalence in Blacks and Mulattos was similar.
The difference in racial prevalence observed
in this work cannot be explained based on age composition of groups, but
it could be casual (despite significant), mainly due to the low prevalence
observed in the White sample (1/148 - 0.7%).
When confronting the anthropological classification
with the genetic composition, a marked process of ethnic admixture is
observed in that region. What strongly suggests that Bantu ancestry is
associated to a higher prevalence of prostate cancer is that in tumor
bearers the proportion of African alleles was 49% whereas in those without
cancer it was 31.7% (Table-1). However, it is remarkable that the proportion
of Amerindian alleles in volunteers without cancer was approximately 18
times higher (9.2%) than the one observed in neoplasia bearers (0.5%).
Could it be that the Amerindian (Asiatic) genes were counterbalancing
the role of African genes in a higher predisposition to cancer?
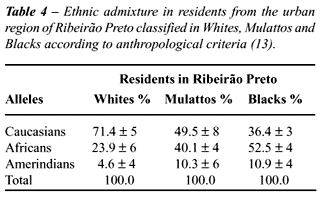
The explanation for the difficulty in demonstrating
a higher prevalence of tumor in Southeastern Blacks is not simple with
data published up to now. One study about miscegenation in a Southeastern
region using the same criteria for anthropological (considering also the
ancestry) and genetic (VNTR) classification, as it can be seen on Table-4
(13), seems to show some ethnic differences between Southeastern and Northeastern
samples. The sample studied in the Northeast suggests a markedly lower
crossing proportion between Caucasian and Indians since the percentage
of Amerindian alleles in Whites was approximately 2.5 times lower (4.6%)
than in the Northeastern sample (11.7%). That means that the proportion
of Amerindian alleles in the Northeastern Whites sample is 154% higher
than the Southeastern one, which would allow us to speculate if this would
be the factor responsible for the low prevalence of tumor in Whites of
that city. Other differences are less significant, even though it can
be stressed that in the Southeast the proportion of African alleles in
Whites was 14.9% higher, of Amerindian alleles in Blacks was 23.8% higher
and of African alleles in Blacks was 14.3% higher than those found in
the Northeastern. Thus, it is possible that the difference in prevalence
among regions can be explained by ethnic diversity. Another explanation
could be the lower longevity of Blacks pointed out in one of the Southeastern
studies (7). However, maybe the main reason is the difference in criteria
for anthropological classification, or even the difficulty or limitations
for its application in a mixed population, because it is not clear if
in other screenings the racial antecedent was considered.
Our study also suggests that the sensitivity
of tPSA, cut-off of 4 ng/ml, in the group of volunteers with tPSA below
or equal to 10 ng/ml, is unsatisfactory. Of the 13 tumors found in this
range, 6 were associated to tPSA < 4 ng/ml, and 4 (30.7%) of them would
not have been diagnosed because the digital rectal examination did not
suggest a tumor. On the other hand, the cut-off of 2 ng/ml presented a
very low specificity, which meant a large proportion of unnecessary biopsies.
The combination of tPSA cut-off in 2.5 ng/ml associated to additional
criteria for indication of biopsy, such as free/total PSA ratio in 20%
or PSAD in 0.08, seemed to represent a better option than employing only
tPSA in all other cut-off levels. In the literature, results are controversial
and there is no agreement about the best cut-off level for tPSA, as well
as about the benefits gained with the associated use of the free/total
PSA ratio and tPSA density, but one can note a tendency towards recommending
a reduction in the cut-off level of tPSA and applying associated parameters
in order to improve the test specificity (4,6). Maybe the most convincing
recent multiracial study about the subject involving the screening and
the follow-up of 12,902 men, of whom 7,541 were considered without prostate
disease, showed that 95% of the sample without disease had tPSA £
2.45 ng/ml, and this is the reason why this value was proposed as the
upper limit for normality (19). Nevertheless, it must be stressed that
the cut-off level of the free/total PSA coefficient that is able to maintain
the tPSA sensitivity above 90%, in the same sample of patients, varies
considerably with the brand of reagent that is used (20).
CONCLUSION
The prevalence of prostate cancer in Ipirá, Bahia, Brazil, was 5.7%, being higher in Blacks and Mulattos than in Whites. In cancer bearers the mean proportion of African alleles was 49%, whereas in those without cancer it was 31.7%. In volunteers with tPSA £ 10 ng/ml, the use of a cut-off level for tPSA in 2.5 ng/ml, the free/total PSA fraction of 20% and tPSA density of 0.08, represented a better option for indication of biopsies than the isolated use of tPSA in a cut-off level of 4 ng/ml.
_____________________________________
The project was supported by the Research
Support Foundation of the State of São Paulo
(FAPESP) and by the Teaching, Research
and Assistance Support Foundation of the
General Hospital of Ribeirão Preto – USP(FAEPA).
REFERENCES
- Oesterling JE, Jacobsen SJ, Chute CG, Guess HA, Girman CJ, Panser LA, et al.: Serum PSA in a community-based population of healthy men: establishment of age-specific reference range. JAMA 1993; 270: 860-4.
- Kronkad A, Lai H, Lamm SH, Lai S: Mortality in prostate cancer. J Urol. 1996; 156: 1084-91.
- Myrtle JF, Klimley PG, Ivor IP, Bruni JF: Clinical utility of prostate specific antigen (PSA) in the management of prostate cancer. Hybritec Inc., San Diego, CA, 1986: 1-4.
- Siegal J, Brawer MK: Prostate Specific Antigen. In: Kaisary AV, Murphy GP, Denis L, Griffiths K (ed.), Text Book of Prostate Cancer, Pathology, Diagnosis and Treatment. London, Martin Dunitz. 1999; pp. 121-41.
- Gann PJ, Hennekens CH, Stampfer MJ: A prospective evaluation of plasma prostate specific antigen for detection of prostatic cancer. JAMA 1995; 273: 289-94.
- Arcangeli CG, Ornstein DK, Keeth DW, Andriole GL: Prostate-specific antigen as a screening test for prostate cancer. The United States experience. Urol Clin North Am. 1997; 24: 299-306.
- Martins AC, Reis RB, Suaid HJ, Maciel LM, Cologna AJ, Falconi RA: Screening for carcinoma of the prostate in volunteers. Int Braz J Urol. 2000; 26: 516-22.
- Antonopoulos IM, Pompeo AC, El Hayek OR, Sarkis AS, Alfer Jr W, Arap S: Results of prostate cancer screening in non-symptomatic men. Int Braz J Urol. 2001; 27: 227-34.
- Glina S, Toscano Jr IL, Mello LF, Martins FG, Vieira VL, Damas CG: Results of screening for prostate cancer in a community hospital. Int Braz J Urol. 2001; 27: 235-43.
- Curtin, PD: The atlantic slave trade: a census. Madson/London, University of Wisconsin Press, 1969.
- Zago MA, Figueiredo MS, Ogo SH: Bantu bs cluster haplotype predominates among Brazilian blacks. Am J Phys Anthropol. 1992; 88: 295-8.
- Antonarakis SE, Boehm CD, Serjeant GR, Theisen CE, Dover GJ, Kazazian Jr HH: Origin of the bs - globin gene in blacks: The contribution of recurrent mutation or gene conversion or both. Proc Natl Acad Sci USA. 1984; 81: 3853-6.
- Silva Jr, WA: Genetic Diversity in Afro-Brazilian Populations. Thesis. Ribeirão Preto School of Medicine - USP, 1999 [in Portuguese].
- Bortolini MC, da Silva-Junior WA, Weimer TA, Zago MA, de Guerra DC, Schneider MP, et al.: Protein and hypervariable tandem repeat diversity in eight African-derived South American populations: inferred relationships do not coincide. Hum Biol. 1998; 70: 443-61.
- Eskew LA, Bare LR, McCullough DL: Systematic 5 region prostate biopsy is superior to sextant method for diagnosing carcinoma of the prostate. J Urol. 1997; 157: 199-203.
- Lewis PO, Zaykin D: Genetic Data Analysis: computer program for the analysis of allelic data. Version 1.0, Internet, GDA Home Page <http://chee.unm.edu/gda/>, 1997.
- Chakraborty R: Gene admixture in human populations: Models and predictions. Am J Phys Anthropol. 1986; 29: 1-5.
- Fonseca FP, Veneziano DB, Betti RC, Okawa CO: Serum levels of prostate specific antigen in patients screened for prostate cancer. Int Braz J Urol. 2001; 27: 32-6.
- Cheli CD, Levine R, Cambetas DR, Kolker JD, Roberts SB: Age-related reference ranges for complexed prostate-specific antigen and complexed/total prostate-specific antigen ratio: results from East Texas Medical Center Cancer Institute screening campaign. Urology 2002; 60 (Suppl A): 53-9.
- Taille A, Houlgatte A, Houldellete P, Goluboff ET, Berlizot P, Ricordel I: Influence of free-to-total prostate specific antigen variability on the early diagnosis of prostate cancer: a comparative study of three immunoassays. Br J Urol. 1998; 82: 389-92.
___________________
Received: April 4, 2003
Accepted after revision: May 20, 2003
_______________________
Correspondence address:
Dr. Antonio C. P. Martins
Hospital das Clínicas da FMRP - USP
Av. Bandeirantes, 3900 / 9º andar
Ribeirão Preto, SP, 14049-900, Brazil
Fax: + 55 16 633-0836
E-mail: acpmartins@convex.com.br